SIR Epidemic Model
Compartments, Thresholds, and Numerical Simulation
The SIR model is a classical compartmental model in epidemiology (Kermack and McKendrick 1927; Keeling and Rohani 2008). It is also a good reminder that the solve_ivp workflow applies to small systems just as naturally as to scalar equations.
We split the population into:
- \(S(t)\): susceptible
- \(I(t)\): infected
- \(R(t)\): recovered
and describes how individuals move between these compartments over time.
We will use the normalized SIR model (fractions of the population, so \(S+I+R=1\)):
\[ \begin{aligned} \dot S &= -\beta SI,\\ \dot I &= \beta SI - \gamma I,\\ \dot R &= \gamma I. \end{aligned} \tag{1}\]
Here \(\beta\) controls transmission and \(\gamma\) controls recovery.
Conservation of Total Population
For the normalized model,
\[ S(t) + I(t) + R(t) = 1. \]
That means the full state has three components, but only two are independent. The model still fits the same initial value problem structure as the scalar examples.
Implement the Right-Hand Side
The numerical solver only needs one function that returns the three derivatives.
def sir_model(t, y, beta, gamma):
s, i, r = y
dsdt = -beta * s * i
didt = beta * s * i - gamma * i
drdt = gamma * i
return [dsdt, didt, drdt]A Useful Threshold
The ratio
\[ R_0 = \frac{\beta}{\gamma} \]
is the basic reproduction number in the normalized model. When \(R_0 > 1\) and the susceptible fraction is still large, the infected compartment initially grows. When \(R_0 < 1\), infection decays instead of taking off.
Reference Implementation
A working implementation is provided in:
amlab/odes_1d/sir_model.py
The model function is sir_model(t, y, beta, gamma) and the plotting helper is plot_sir_model(...).
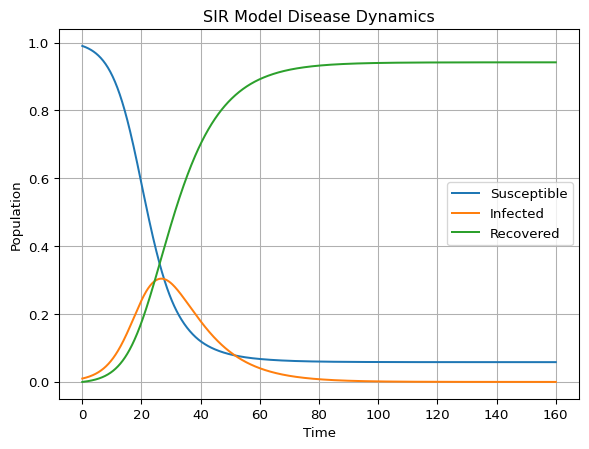
Render-time Figure
The figure below is generated at render-time from the reference script.

Simulate with solve_ivp()
The solver call follows the same pattern as every other ODE example in the course:
solution = solve_ivp(
sir_model,
t_span=(0, 160),
y0=[0.99, 0.01, 0.0],
args=(0.3, 0.1),
t_eval=np.linspace(0, 160, 1000),
)What to Look For
- The susceptible population decreases monotonically.
- The infected population typically rises, peaks, and then falls.
- The recovered population grows monotonically.
- Changing \(\beta\) mostly affects how fast the outbreak grows.
- Changing \(\gamma\) mostly affects how quickly the outbreak resolves.
Exploration
- Increase \(\beta\) while keeping \(\gamma\) fixed. What happens to the peak of \(I(t)\)?
- Increase \(\gamma\) while keeping \(\beta\) fixed. Does the epidemic end sooner?
- Try different initial infected fractions \(I(0)\). Do you always see an outbreak?
Run Locally
To run the standalone script and show the plot:
python amlab/odes_1d/sir_model.pyWhat’s Next?
The SIR model shows how the same solver handles several coupled state variables. The next page returns to a scalar nonlinear rate law with saturation.