import matplotlib.pyplot as plt
import networkx as nx
SEED = 7
def summarize_graph(G, name="G"):
"""Print a small summary table of common network statistics."""
n = G.number_of_nodes()
m = G.number_of_edges()
avg_k = 2 * m / n if n > 0 else 0
density = nx.density(G)
# Connectivity (undirected)
is_connected = nx.is_connected(G) if n > 0 else False
n_components = nx.number_connected_components(G) if n > 0 else 0
# Work on largest connected component when disconnected
if n == 0:
L = float("nan")
C = float("nan")
else:
C = nx.average_clustering(G)
if is_connected:
L = nx.average_shortest_path_length(G)
else:
largest_cc = max(nx.connected_components(G), key=len)
H = G.subgraph(largest_cc).copy()
L = nx.average_shortest_path_length(H)
print(f"--- {name} ---")
print(f"N = {n}, E = {m}, <k> = {avg_k:.2f}, density = {density:.4f}")
print(f"connected = {is_connected}, components = {n_components}")
print(f"clustering C = {C:.3f}")
print(f"avg shortest path L = {L:.3f} (largest CC if disconnected)")
def plot_degree_histogram(G, title="Degree distribution"):
degrees = [deg for _, deg in G.degree()]
plt.figure(figsize=(5, 3))
plt.hist(degrees, bins=range(min(degrees), max(degrees) + 2), align="left", rwidth=0.9)
plt.xlabel("Degree k")
plt.ylabel("Count")
plt.title(title)
plt.tight_layout()
plt.show()Networks: Graph Models
Random, Small-World, Scale-Free
Real networks (social, transportation, communication) are not arbitrary: they tend to have short paths, clustering, and sometimes hubs (Newman 2018).
In this page, we introduce three classic synthetic graph models that help us reproduce (some of) these properties:
- Random graphs (Erdős–Rényi / Gilbert)
- Small-world graphs (Watts–Strogatz)
- Scale-free graphs (Barabási–Albert preferential attachment)
We will also compute a few summary metrics for each model:
- Average degree \(\langle k \rangle\)
- Density \(D\)
- Average clustering coefficient \(C\)
- Average shortest path length \(L\) (on the largest connected component if needed)
If you have not seen these metrics before, the definitions and examples are in the previous page: Network Connectivity.
Random graphs (Erdos–Rényi / Gilbert)
In the Gilbert model \(G(N, p)\), we consider every possible pair of nodes and include the edge independently with probability \(p\) (Gilbert 1959; Erdos and Renyi 1959).
Key properties (typical, for large \(N\)):
- Degree distribution is approximately binomial, and approaches Poisson in the sparse limit.
- Average clustering is approximately \(C \approx p\).
- Typical path lengths scale like \(L \sim \log N\) when the graph is above the connectivity threshold.
In NetworkX, you can create this model with nx.erdos_renyi_graph(n, p) (alias: nx.gnp_random_graph).
- Select a pair of nodes, say i and j.
- Generate a random number r between 0 and 1. If r < p, then add a link between i and j.
- Repeat (1) and (2) for all pairs of nodes.
# Random graph G(N, p)
G_random = nx.erdos_renyi_graph(n=80, p=0.06, seed=SEED)
pos = nx.spring_layout(G_random, seed=SEED)
nx.draw(G_random, pos=pos, node_size=120, node_color="lightgray", edge_color="gray")
plt.title("Erdos–Rényi random graph")
plt.show()
summarize_graph(G_random, name="Random (G(N,p))")
plot_degree_histogram(G_random, title="Random graph: degree distribution")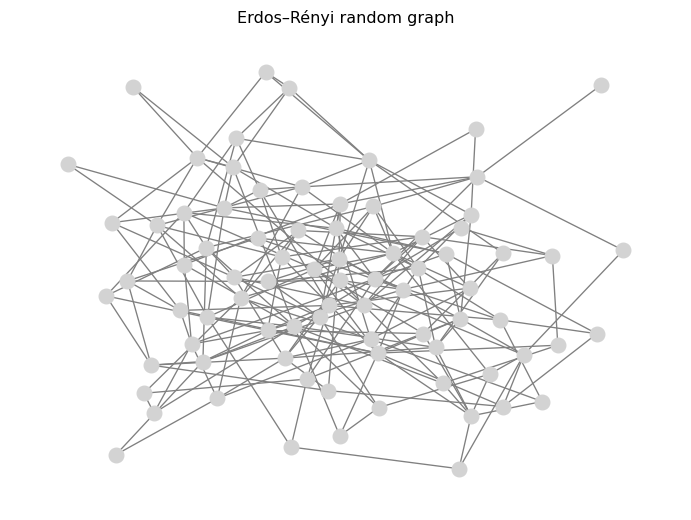
--- Random (G(N,p)) ---
N = 80, E = 211, <k> = 5.28, density = 0.0668
connected = True, components = 1
clustering C = 0.078
avg shortest path L = 2.766 (largest CC if disconnected)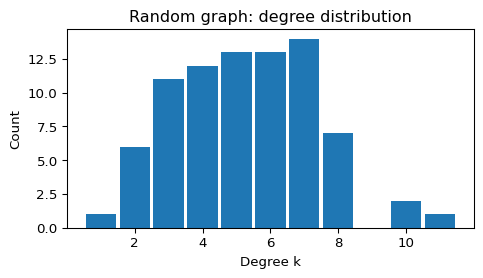
Another popular variant is \(G(N, m)\), where we fix the number of edges \(m\) instead of an edge probability. In NetworkX:
nx.gnm_random_graph(n, m)implements \(G(N,m)\)nx.erdos_renyi_graph(n, p)implements \(G(N,p)\)
# Random graph G(N, m)
G_gnm = nx.gnm_random_graph(n=80, m=180, seed=SEED)
summarize_graph(G_gnm, name="Random (G(N,m))")
plot_degree_histogram(G_gnm, title="G(N,m): degree distribution")--- Random (G(N,m)) ---
N = 80, E = 180, <k> = 4.50, density = 0.0570
connected = False, components = 2
clustering C = 0.039
avg shortest path L = 3.026 (largest CC if disconnected)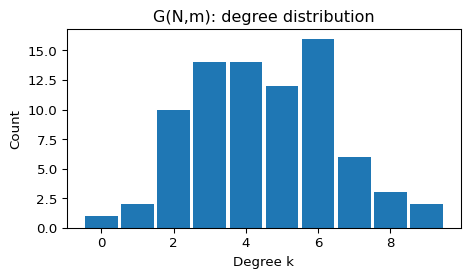
Small-world graphs (Watts–Strogatz)
Watts–Strogatz networks start from a ring lattice (high clustering, long paths) and then rewire edges with probability \(p\) (Watts and Strogatz 1998).
Key properties:
- For small \(p\) (e.g. \(p \in [0.01, 0.2]\)), you often get high clustering and short path lengths (“small-world”).
- Degree distribution stays relatively narrow (most nodes have degree near \(k\)).
In NetworkX:
nx.watts_strogatz_graph(n, k, p)generates the classic model.kis the number of neighbors of each node in the initial ring (typically even).
- Begin with a ring of \(N\) nodes
- Connect each node to its \(k\) nearest neighbors (or \(k-1\) if k is odd).
- For each edge \((u, v)\), with probability \(p\), replace edge \((u, v)\) with \((u, w)\) where \(w\) is not a neighbor of \(u\).
# Watts-Strogatz small-world graph with n nodes, k neighbors, and probability p of rewiring
G_sw = nx.watts_strogatz_graph(n=80, k=6, p=0.08, seed=SEED)
pos = nx.spring_layout(G_sw, seed=SEED)
nx.draw(G_sw, pos=pos, node_size=120, node_color="lightblue", edge_color="gray")
plt.title("Watts–Strogatz small-world graph")
plt.show()
summarize_graph(G_sw, name="Small-world (Watts–Strogatz)")
plot_degree_histogram(G_sw, title="Small-world: degree distribution")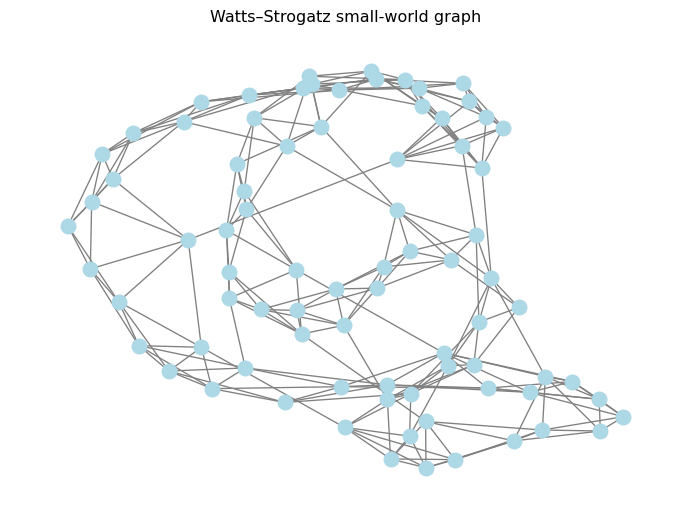
--- Small-world (Watts–Strogatz) ---
N = 80, E = 240, <k> = 6.00, density = 0.0759
connected = True, components = 1
clustering C = 0.494
avg shortest path L = 3.694 (largest CC if disconnected)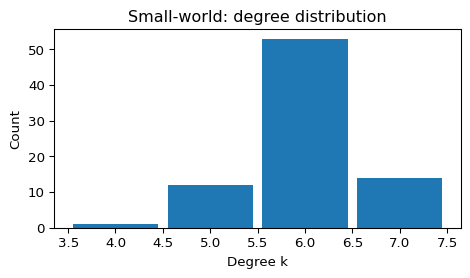
Scale-free graphs (preferential attachment)
Many real networks exhibit a heavy-tailed (sometimes approximately power-law) degree distribution: most nodes have small degree, but a few nodes become hubs (Barabasi and Albert 1999; Newman 2018).
The Barabási–Albert (BA) model generates this effect via preferential attachment (Barabasi and Albert 1999):
- Nodes that already have many edges are more likely to receive new edges.
Key properties (typical):
- Degree distribution is heavy-tailed (often summarized as “scale-free”).
- Small average path length (hubs create shortcuts).
- Clustering is usually lower than small-world models (but there are variants that increase clustering).
In NetworkX:
nx.barabasi_albert_graph(n, m)where each new node attaches tomexisting nodes.
- Start with a clique of \(m + 1\) nodes.
- Select \(m\) different nodes at random, weighted by their degree.
- Add a new node \(i\) and link it with the \(m\) nodes from the previous step.
- Repeat 2-3 until there are N nodes in the graph.
# Barabasi-Albert preferential attachment graph with n nodes and m edges
G_sf = nx.barabasi_albert_graph(n=200, m=2, seed=SEED)
pos = nx.spring_layout(G_sf, seed=SEED)
nx.draw(G_sf, pos=pos, node_size=35, node_color="lightgreen", edge_color="gray", alpha=0.8)
plt.title("Barabási–Albert (scale-free-ish) graph")
plt.show()
summarize_graph(G_sf, name="Scale-free (Barabási–Albert)")
plot_degree_histogram(G_sf, title="Scale-free: degree distribution")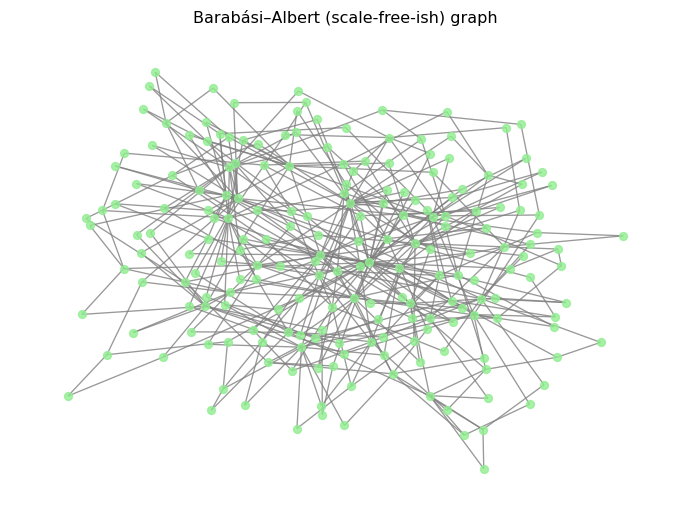
--- Scale-free (Barabási–Albert) ---
N = 200, E = 396, <k> = 3.96, density = 0.0199
connected = True, components = 1
clustering C = 0.075
avg shortest path L = 3.483 (largest CC if disconnected)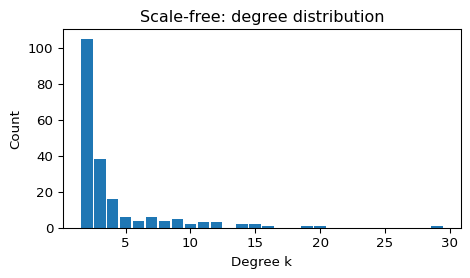
The BA model is the most common introduction to “scale-free” networks. NetworkX also provides other generators (some return directed multigraphs), for example:
nx.scale_free_graph(n, seed=...)nx.powerlaw_cluster_graph(n, m, p, seed=...)(adds triangle-closing for higher clustering)
For this course, nx.barabasi_albert_graph is usually the best starting point.
What’s Next?
Ready to simulate epidemics? Continue to: